How to see cells move in a turbulent watery world
Every second, everywhere, we pick up information about the environment around us through senses such as sight, touch, and hearing.
Likewise, trillions of cells inside our bodies are constantly exploring the highly dynamic microenvironment surrounding them. White blood cells, for example, travel through blood vessels and different tissues to fight pathogens to protect us against infection.
To fulfil their biological functions, these cells must have developed ways of sensing and reacting to cues presented in their turbulent, watery, and dark environment.
The question is how.
How do different groups of cells know what environment they are in and decide where to move?
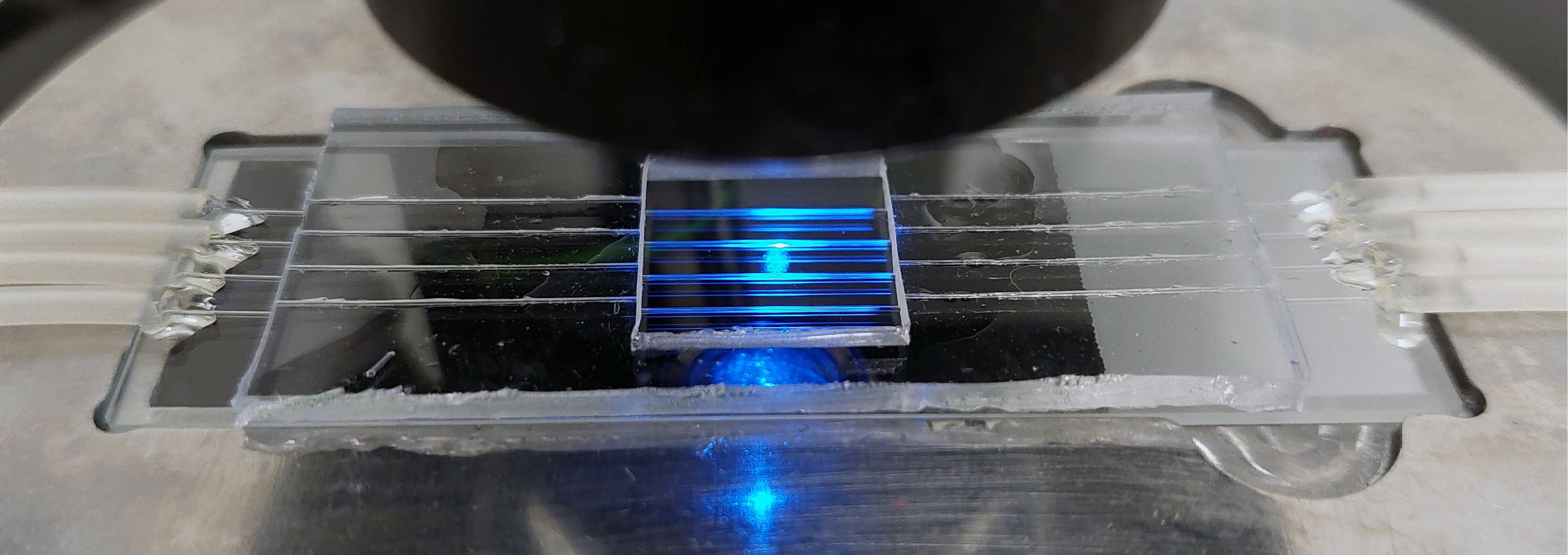

This specialised chip will help researchers approach the answer to this big question. Image: Lee Group
Seeing living cells migrate under pressure
In the 1970s, British cell biologist Michael Abercrombie first scrutinised how cells called fibroblasts move, resulting in several seminal publications on cell migration.
His pioneering work was bolstered by the use of time-lapse photography, a cutting-edge imaging technology at his time.
Abercrombie’s innovative combination of optical technology and biology underpinned the discipline of cell migration, which has since impacted many fields to explore the cellular basis of how wounds heal and how cancer cells metastasise.
However, few studies investigated microenvironment cues, as most imaging experiments were done on simple 2D glass dishes under stagnant liquid conditions.
“But cells live within incredibly challenging watery microscopic environments,” said Dr Woei Ming (Steve) Lee, Head of the Optical Biofluidic Group.
“From mucous in our gut to plasma in our vessels, high levels of viscous, maple syrup-like fluids constantly flow and exert mechanical pressure on cells.”
Therefore, when studying how cells sense and navigate their environments in a living organism, one cannot ignore the effects of fluid flows on cells.
Video: Lee Group
At the John Curtin School of Medical Research, the Lee Group focuses on re-creating cues in the native environment around cells in a controlled laboratory setting and objectively assessing how each cell changes behaviour over several hours to days.
The overarching question the team is trying to answer is whether chaotic fluid flow in a diseased organ alters cell behaviour.
“We want to observe, in the most native state, how different groups of cells respond to fluid-induced mechanical stresses,” said Steve.
But reproducing a disease-related fluid flow state in the lab for observation is easier said than done—commercially available tools are not developed to meet this need.
“We are thus ‘forced’ to invent new tools,” said Steve.
Merging disciplines to solve problems
Having been through diverse training in engineering, physical science and biomedical imaging, Steve enjoys inventing biophysical tools to solve biomedical problems.
His team members share his experience and passion. Importantly, interdisciplinary and diverse training enables the team to jump out of the box whenever needed.
“To us the challenge is two-faceted,” said Dr Yean Jin (Daniel) Lim, a Senior Imaging Scientist working in the Lee Group, “It’s both about recreating the diseased microenvironment we try to study in the lab and seeing how cells respond to that environment without too much perturbation to the cell’s activity”
To address the first problem, Daniel spearheaded the work of transforming a standard multiphoton imaging microscope into a cutting-edge ‘3D printer’.
Inspired by Inner painting—an ancient art form—the team used the microscope to sculpt the inner surfaces of transparent glass capillaries.
This new technique, known as Multiphoton Inner Laser Lithography (MILL), allowed them to create micro-patterned flow channels that recapitulated true-to-life tissue topology and 3D fluid flows with various levels of fluid shear.
“A tool is not necessarily limited by the application that was prescribed to it.” Daniel stressed.
The MILL technique has led to a standardised workflow, where different MILL chips are made to recapitulate different fluidic settings, ranging from blood vessels to lymphatic to stroma tissues.
MILL chip that recapitulates fluidic settings of vascular stenosis. Video: Lee Group
But that only overcame half of the challenges. To observe cell behaviour with minimum perturbation, the team must further innovate how cells are imaged.
“With the ability to reproduce in vivo fluid flow in minutes, we now face the challenge of observing and assessing the biological activities of cells, in their most native context, from millimetre level to nanoscale, literally on the same day,” he said.
Putting cell imaging back into the natural context
Almost all cells in our body navigate in virtual darkness, but at the same time, almost all existing imaging tools need a lot of light to see inside a cell.
“Imagine standing in a desert under continuous sunlight. Light can kill. Cells in your body are not used to seeing lots of light, so they are sensitive to different light levels,” Steve explained.
Conventional lasers that consist of highly intense light can cause a cell-damaging process known as phototoxicity, killing cells indirectly.
As the researchers want to see cell migration in its most natural state, they need to image the cell with as little light as possible.
“Thankfully, some of the assumptions previously perceived as ‘rules’ of light microscopy can be bent,” said Tienan Xu, PhD Student at the Lee Group.
For instance, phototoxicity arises from imaging a cell with intense light multiple times over several minutes to hours. But the team found that taking many images is not essential for recording 3D information. It is possible to tune the light and to design a camera that can read instantaneously where each photon arrives from in 3D and in a single snapshot.
Xu recently published a computational tool that can achieve 3D imaging volume using a single 2D image, which frees the cells from being exposed to highly intense light many times and thus substantially reduces phototoxicity. The team has filed the technology for a patent, with a few industry partners expressing interest.
Meanwhile, other lab members managed to open ways to acquire good-quality quantitative data in live cell imaging with less light.
Previous PhD student Yujie Zheng (now working at BGI Genomics), for instance, developed a new form of nanoscopy to see the individual organelles of each migrating cell.
High resolution imaging of endothelial cell. Video: Lee Group
Graduating PhD student Avinash Upadhya, who has started a position at the University of Adelaide, on the other hand, developed a programmable microscope using digital optics technologies to record live events of migrating cells in 3D.
Together, these imaging innovations paved the way for a unique “adaptive imaging” workflow, where the imaging tools and microfabrication tools adapt to the targeted biological system rather than the other way around.
“Our inventions arise from pushing the limits of questioning how—how living cells move in a dynamic, in-vivo fluid microenvironment,” said Steve.
As part of the Australian Cancer Research Foundation Centre for Intravital Imaging of Niches for Cancer Immune Therapy (INCITE), the team aims to extend the unique “adaptive imaging” innovations into the pharmacological sector.
“Imagine a world where we can take a wide range of living cancer cells, for example, put them through an assay, and effectively measure their metastatic potential,” remarked Steve, “That is the primary goal, but we can do so much more.”
Innovating tools through motivating minds
While the Lee Group is excited that they are ready to push the boundaries of the field of cell migration, there will be a time when they have to build new useful tools again.
“You cannot innovate in the field if you do not innovate the instruments,” said Daniel, “It is a cycle.”
What fuels the cycle of innovation is new ideas and new minds.

Steve Lee teaching in the class. Image: Lee Group
“Our motto is ‘learn by doing, share by teaching’,” said Steve.
He has started incorporating the lesson in the lab into classrooms by teaching a new undergraduate course named Biomedical Imaging: Principles, Technologies and Applications in Health Sciences.
In the course, students apply broad concepts of biomedical imaging and debate how modern imaging technologies drive medical breakthroughs. Steve hopes this will excite more talented minds to illuminate the future of the field.
“It’s a great time for exploring new biology,” he said, “We learn to see a new world unveiled before us by finding tools to open new eyes.”
*Acknowledge fundings from: The Australian Research Council (ARC), National Health and Medical Research Council (NHMRC), Australian Cancer Research Foundation, AusIndustry, The Australian National University, Discovery Translational Grant.
